library(gapminder)Exercises
Data
Load the gapminder data:
Exercises
Exercise 1: Parameterized plots with ggplot2
- Create a variable to define a year in the
gapminderdata that you want to plot.
Solution
choose_year <- 1952- Edit the code below to use this new year variable.
library(dplyr)
library(ggplot2)
# Median life expectancy per continent
gapminder_1952 <- gapminder |>
filter(year == 1952) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
ggplot(data = gapminder_1952) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
)
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = "Life Expectancy in 1952"
)Solution
library(dplyr)
library(ggplot2)
library(glue)
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
)
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)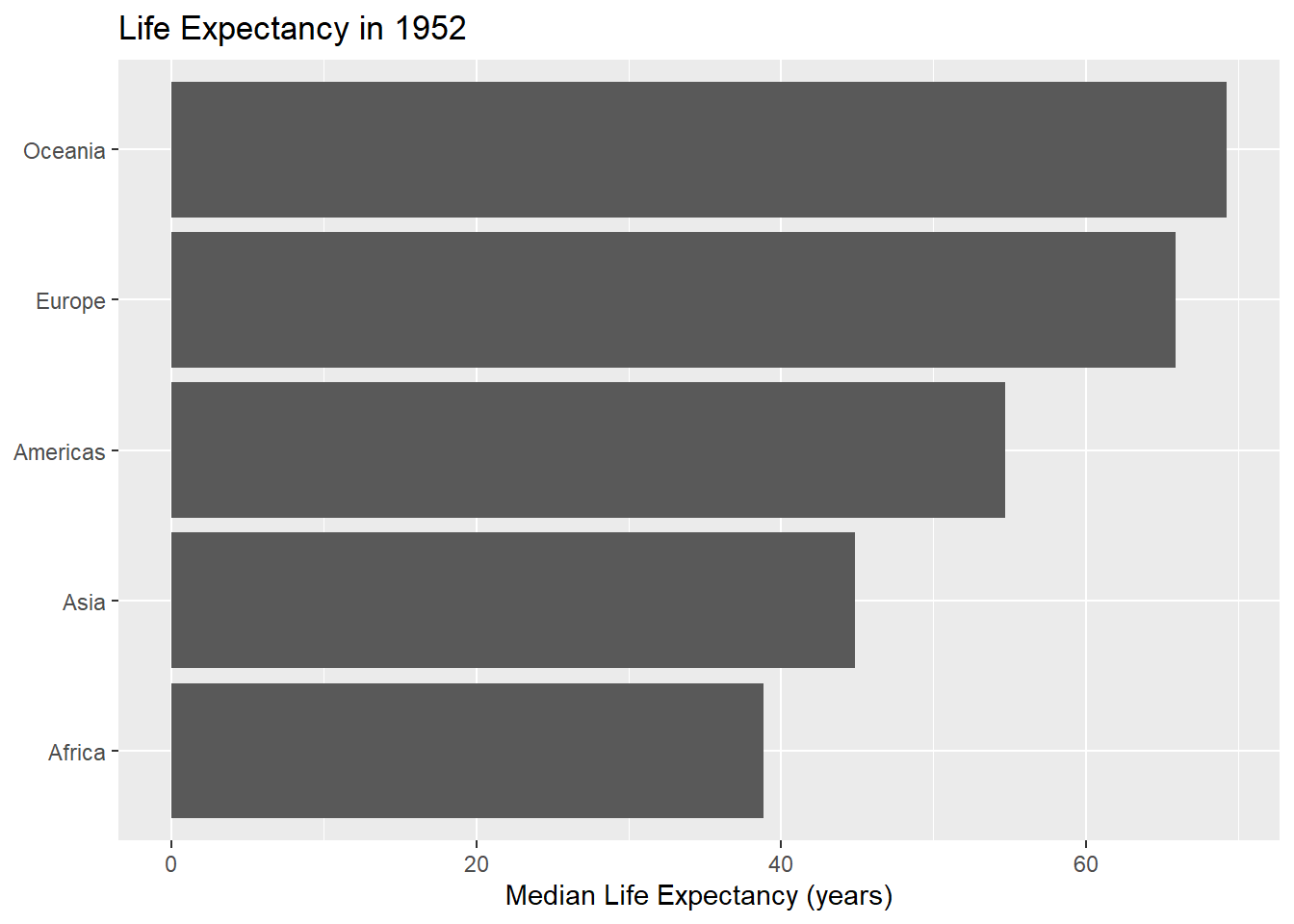
- Turn your code into a function that takes the data and year variable as arguments.
Solution
barchart_year <- function(choose_year, data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
)
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}Check it works:
barchart_year(2002)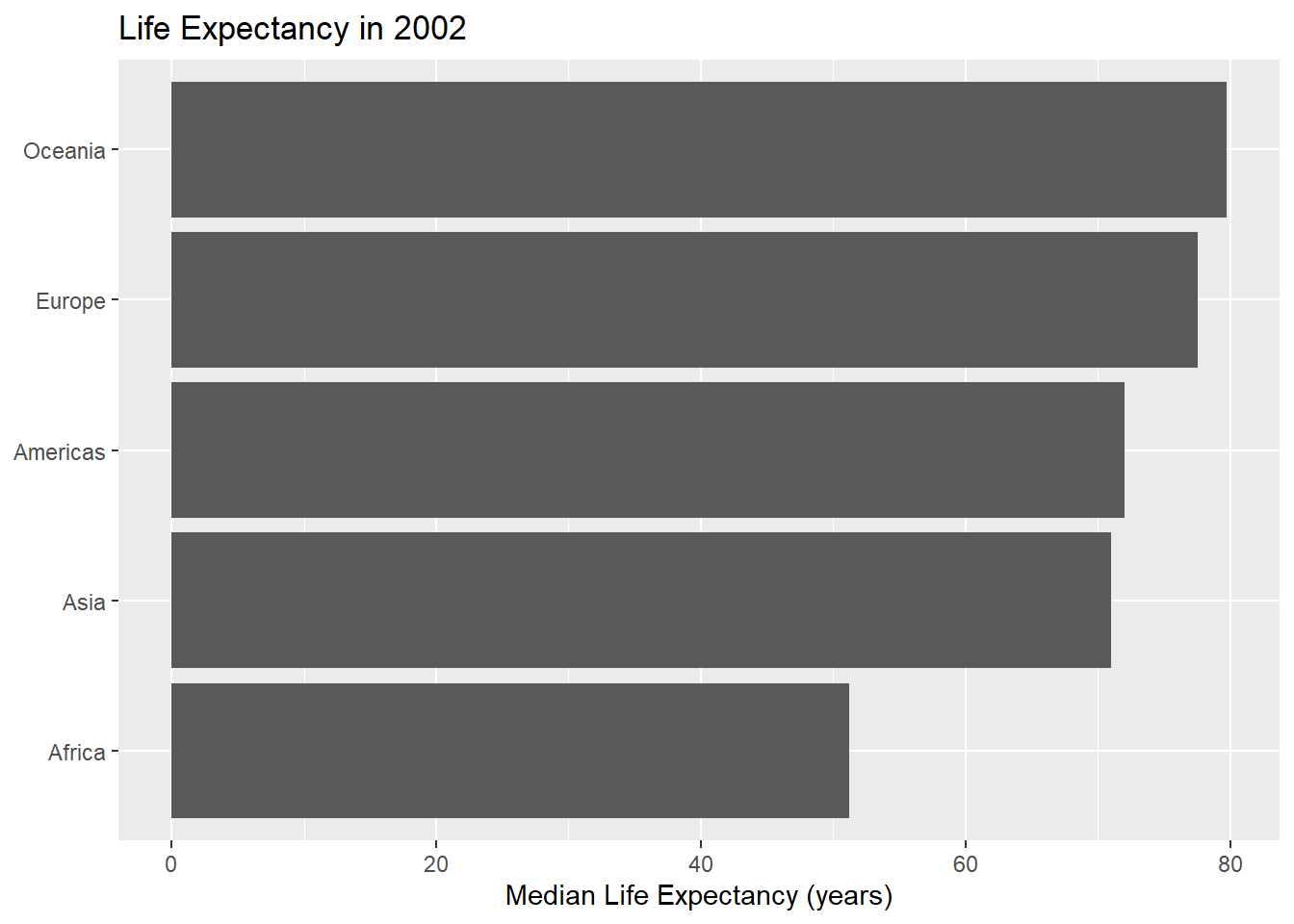
- Bonus: Add another argument to your function that allows a user to define the bar colour.
Solution
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}Check it works:
barchart_year(2002, fill = "purple")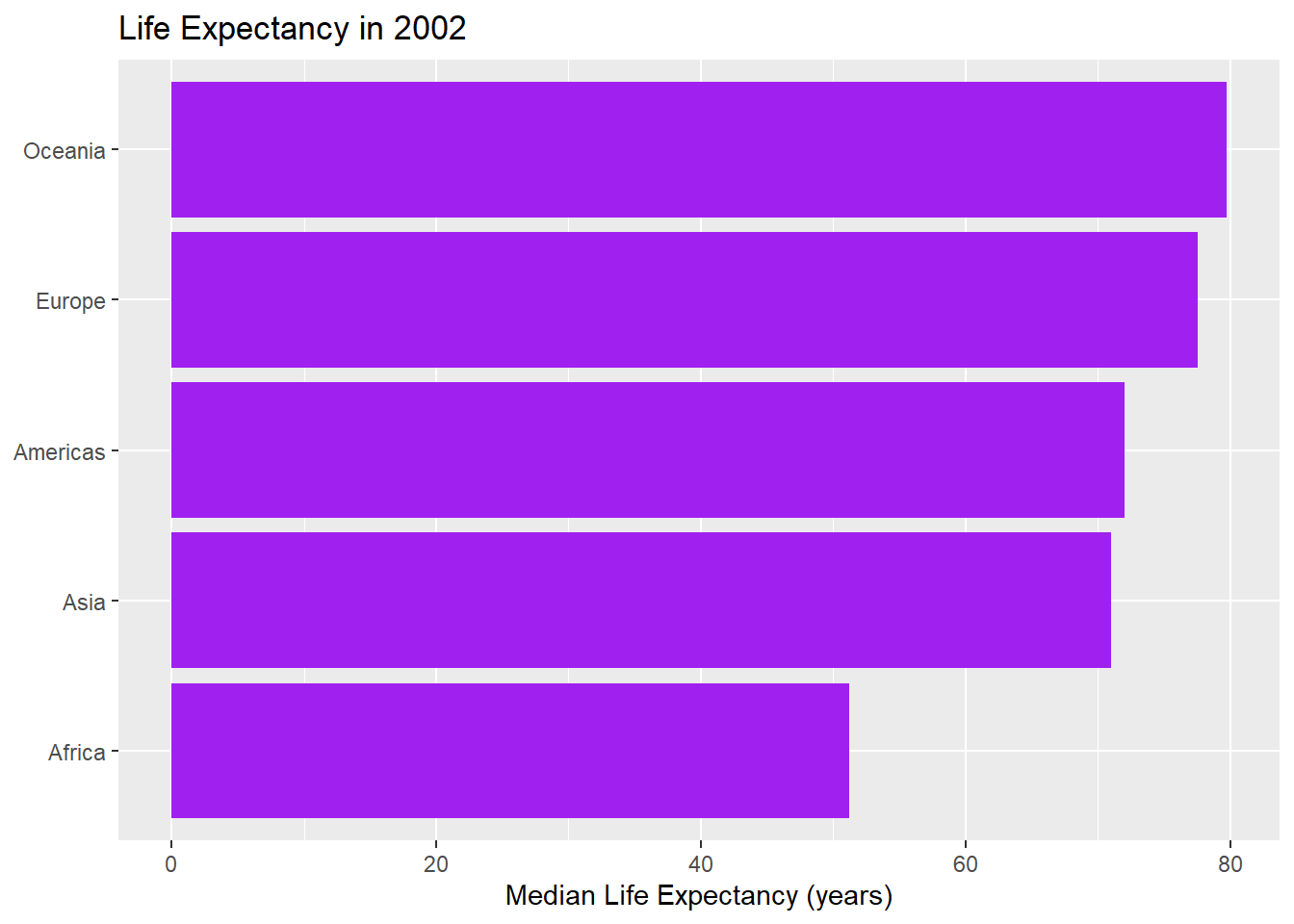
- Bonus: How could you improve your function?
Solution
- Not all year values are included in the data. Add a check that the value is a number that is included in the
gapminderdata. if it isn’t, add a more useful message. - Write some documentation!
Exercise 2: Parameterized reports with Quarto
- Edit the
report.qmdfile to add a parameter for year.
Warningreport.qmd
---
title: "Gapminder Report: 1952"
execute:
echo: false
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year 1952 for each continent.
```{r}
#| label: fig-life-exp
#| fig-cap: "Life expectancy in 1952"
# Edit the code below with your plot function!
plot(1:10, 1:10)
```Solution
---
title: "Gapminder Report: 1952"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year 1952 for each continent.
```{r}
#| label: fig-life-exp
#| fig-cap: "Life expectancy in 1952"
# Edit the code below with your plot function!
plot(1:10, 1:10)
```- Add your plotting function to the quarto document, and pass the parameter into the plotting function. Check it renders.
Solution
---
title: "Gapminder Report: 1952"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year 1952 for each continent.
```{r}
#| label: fig-life-exp
#| fig-cap: "Life expectancy in 1952"
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}
barchart_year(params$choose_year)
```- Change the value of the parameter, and re-render.
Exercise 3: Parameterized text with Quarto
- Edit the sentence
...a plot of the median life expectancy in the year 1952 for each continent...to use inline code to add the document parameter for the year.
Solution
---
title: "Gapminder Report: 1952"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year `r params$choose_year` for each continent.
```{r}
#| label: fig-life-exp
#| fig-cap: "Life expectancy in 1952"
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}
barchart_year(params$choose_year)
```- Use inline code for the title as well.
Solution
---
title: "Gapminder Report: `r params$choose_year`"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year `r params$choose_year` for each continent.
```{r}
#| label: fig-life-exp
#| fig-cap: "Life expectancy in 1952"
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}
barchart_year(params$choose_year)
```- Edit the figure caption to use inline code to add the document parameter for the year.
Solution
---
title: "Gapminder Report: `r params$choose_year`"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year `r params$choose_year` for each continent.
```{r}
#| label: fig-life-exp
#| fig-cap: !expr 'paste("Life expectancy in ", params$choose_year)'
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}
barchart_year(params$choose_year)
```Exercise 4: Rendering multiple reports
- Render a version of your report for the year
1972using thequarto_render()function.
Solution
quarto_render(
input = "report.qmd",
execute_params = list(choose_year = 1972)
)- Render a version of your report for all years in the data.
Solution
walk(
.x = unique(gapminder$year),
.f = ~quarto_render(
input = "report.qmd",
output_file = glue("report_{.x}.html"),
execute_params = list(
choose_year = .x
)
)
)Exercise 5: Conditional content
- Edit the .qmd file from Exercise 3. When the format is HTML show an interactive chart using
plotly, otherwise show a static chart.
Solution
---
title: "Gapminder Report: `r params$choose_year`"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
library(plotly)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year `r params$choose_year` for each continent.
```{r}
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}
plot_output <- barchart_year(params$choose_year)
```
::: {#fig-life-exp fig-cap="Life expectancy in `r params$choose_year`"}
::: {.content-visible when-format="html"}
```{r}
ggplotly(plot_output)
```
:::
::: {.content-visible unless-format="html"}
```{r}
plot_output
```
:::
:::
- When the year parameter is 2007, add a sentence to state that it is the most recent data available.
Solution
---
title: "Gapminder Report: `r params$choose_year`"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
library(plotly)
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year `r params$choose_year` for each continent.
```{r}
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}
plot_output <- barchart_year(params$choose_year)
```
::: {#fig-life-exp fig-cap="Life expectancy in `r params$choose_year`"}
::: {.content-visible when-format="html"}
```{r}
ggplotly(plot_output)
```
:::
::: {.content-visible unless-format="html"}
```{r}
plot_output
```
:::
:::
```{r}
latest_begin <- if (params$choose_year != 2007) "::: {.content-hidden}"
latest_end <- if (params$choose_year != 2007) ":::"
```
`r latest_begin`
This is the most recent data available.
`r latest_end`
- Bonus: Can you edit the logic you applied in question 2, to check programmatically if it’s the last year in the data?
Solution
last_year <- max(gapminder$year)
latest_begin <- if (params$choose_year != last_year) "::: {.content-hidden}"
latest_end <- if (params$choose_year != last_year) ":::"Exercise 6: Parameterized report sections
Separate out the
barchart_year()function into a separate R script.Load it in using
file:instead.
Solution
Create a .R file and put the function in it. You don’t need to make any changes.
setup.R
barchart_year <- function(
choose_year, fill = "blue", data = gapminder) {
# Median life expectancy per continent
gapminder_year <- gapminder |>
filter(year == choose_year) |>
group_by(continent) |>
summarise(
median_lifeExp = median(lifeExp)
)
# Bar chart
g <- ggplot(data = gapminder_year) +
geom_col(
mapping = aes(
x = median_lifeExp,
y = reorder(continent, median_lifeExp)
),
fill = fill
) +
labs(
x = "Median Life Expectancy (years)", y = NULL,
title = glue("Life Expectancy in {choose_year}")
)
return(g)
}The use file: to load it in.
---
title: "Gapminder Report: `r params$choose_year`"
execute:
echo: false
params:
choose_year: 1952
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
```{r}
#| file: setup.R
```
This is a report using the [Gapminder](https://www.gapminder.org/data/) data. @fig-life-exp shows a plot of the median life expectancy in the year `r params$choose_year` for each continent.
::: {.cell}
```{.r .cell-code}
barchart_year(params$choose_year)
```
::: {.cell-output-display}
{#fig-life-exp width=672}
:::
:::
- Create a child file containing the year as a section heading, followed by the chart.
Solution
- Render document with a section for each year in the data.
Solution
---
title: "Gapminder Report
execute:
echo: false
---
```{r}
#| label: load-pkgs
#| warning: false
library(gapminder)
library(dplyr)
library(ggplot2)
library(glue)
```
```{r}
#| file: setup.R
```
```{r}
#| output: asis
all_years <- as.character(unique(gapminder$year))
for (i in seq_len(length(all_years))) {
params <- list(choose_year = all_years[i])
a <- knitr::knit_child('_child.qmd', quiet = TRUE)
cat(a, sep='\n')
}
```